Accession number BBBC038 · Version 1
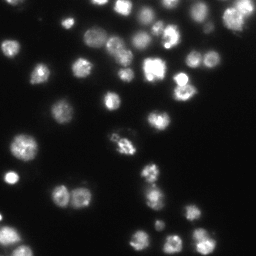
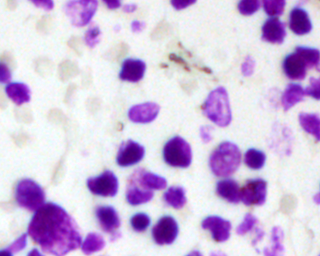
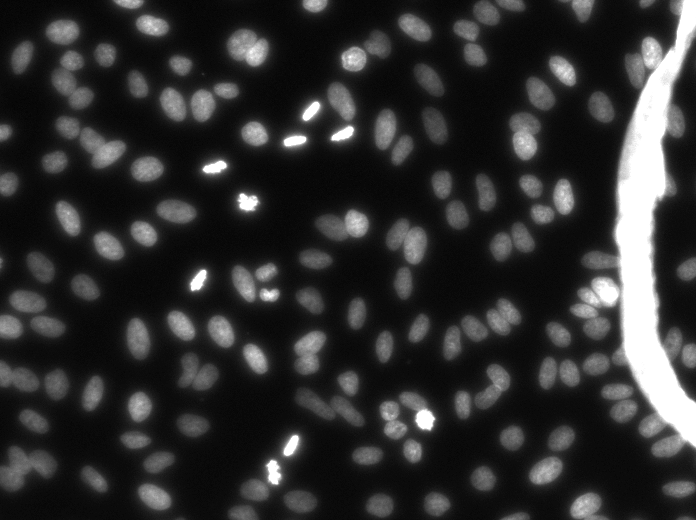
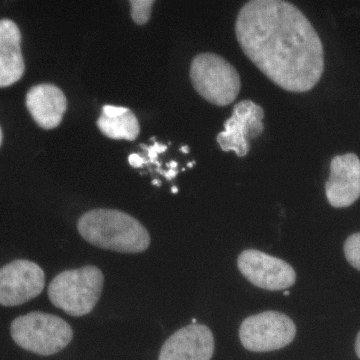
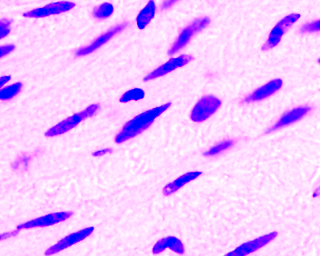
Example images
 |
 |
 |
 |
 |
Description of the biological application
This image data set contains a large number of segmented nuclei images and was created for the Kaggle 2018 Data Science Bowl sponsored by Booz Allen Hamilton with cash prizes. The image set was a testing ground for the application of novel and cutting edge approaches in computer vision and machine learning to the segmentation of the nuclei belonging to cells from a breadth of biological contexts.
Images
These images form a diverse collection of biological images collectively containing tens of thousands of nuclei. The variety within the data set reflects the type of images collected by research biologists at universities, bio-techs, and hospitals. The nuclei in the images are derived from a range of organisms including humans, mice, and flies. In addition, nuclei have been treated and imaged in a variety of conditions including fluorescent and histology stains, several magnifications, and varying quality of illumination. Finally, nuclei appear in different contexts and states including cultured mono-layers, tissues, and embryos, and cell division, genotoxic stress, and differentiation. The dataset is designed to challenge an algorithm's ability to generalize across these variations.
Each image is represented by an associated ImageId. Files belonging to an image are contained in a folder with this ImageId. Within this folder are two subfolders:
- images contains the image file
- masks contains the segmented masks of each nucleus. This folder is only included in the training set. Each mask contains one nucleus. Masks are not allowed to overlap (no pixel belongs to two masks). The second stage dataset contains from experimental conditions not present in the first stage. To deter hand labeling, it also contains images that are ignored in scoring. See the stage2_solution_final.xls file column containing the word "Ignored".
stage2_test_final.zip (289.7 MB)
Ground truth 
In addition to the images there is an accompanying collection of annotations. The annotations were originally created by the Broad Imaging Platform. The annotations take the form of a collection of masks for each image of nuclei. Each mask is a PNG file that contains the segmentation of exactly one nucleus in a folder with the same name as the image it refers to. Like the masks, the images of nuclei are also PNG.
The ground truth and annotations were originally created by the Broad Imaging Platform using a combination of GIMP and a web-based annotation tool created internally.
stage1_train_labels.csv (8.1 MB)
stage2_solution_final.csv (1.5 MB)
These are examples of the submission format:
stage1_sample_submission.csv (5 KB)
stage2_sample_submission_final.csv (208 KB)
Some publicly available improved annotations are available in these two github repositories. We would welcome someone submitting these to BBBC in the same format as a new version:
[Data science bowl 2018 training set improved (Github repo)]
[Kaggle Data Science Bowl 2018 dataset fixes (Github repo)]
For more information
These images were curated from a variety of sources (below) by the Imaging Platform at the Broad Institute for the 2018 Data Science Bowl. Please contact the Imaging Platform with any inquiries.
Contributors
- Riki Eggert, King's College London
- Donna McPhie, McLean Hospital
- Andrew Bradley and Gustavo Carneiro
- Mariko Taga, Columbia University
- Matthew Stachler, Brigham and Women's Hospital
- Chris Lee, MIT
- Alexander Chamessian, Ji Lab, Duke University
- Florian Barthelemy, Miceli Lab, Center for Duchenne muscular dystrophy, UCLA
- Lorraine Montel, Ecole Normale Superieure
- Glyn Nelson, Newcastle University
- Tim Becker, Fraunhofer EMB
- Maria Frias, Foster Lab, Hunter College
- Philipp Keller
- Christian Marinaccio, Northwestern University
- Vasiliy Chernyshev, Skoltech
- several biologists who wished to remain anonymous
- And of course, the Carpenter lab, Broad Institute
Published results using this image set
These datasets will be evaluated in a publication to be submitted.
Recommended citation
"We used image set BBBC038v1, available from the Broad Bioimage Benchmark Collection [Caicedo et al., Nature Methods, 2019]."
Copyright
 Copyright: CC0. To the extent possible under law, the various contributors of the imagesets have waived all copyright and related or neighboring rights to BBBC038v1.
Copyright: CC0. To the extent possible under law, the various contributors of the imagesets have waived all copyright and related or neighboring rights to BBBC038v1.