Accession number BBBC018 · Version 1
Example Images
|
|
|
|
|
|
|
Actin blebs |
Actin dots |
Anaphase/Telephase |
Angular Cell Edges |
Cresent Nuclei |
|
|
|
|
|
|
|
Long Projections |
Metaphase |
Motile |
Peas in a Pod |
Peripheral Actin |
|
|
|
|
|
|
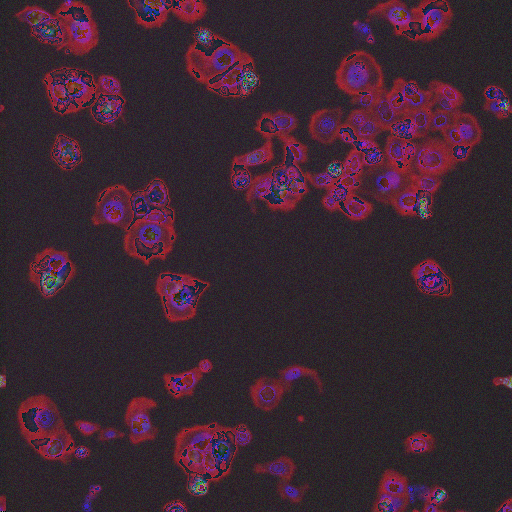
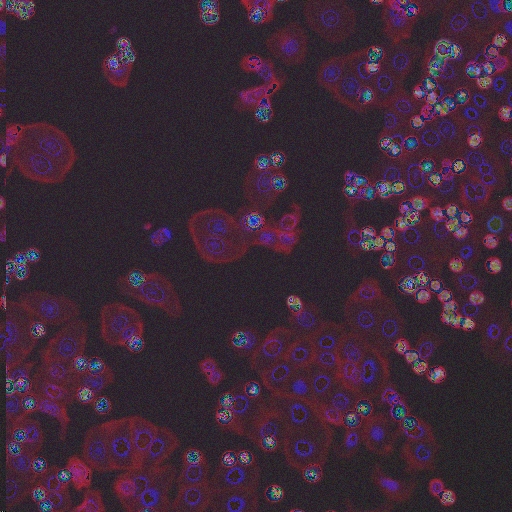
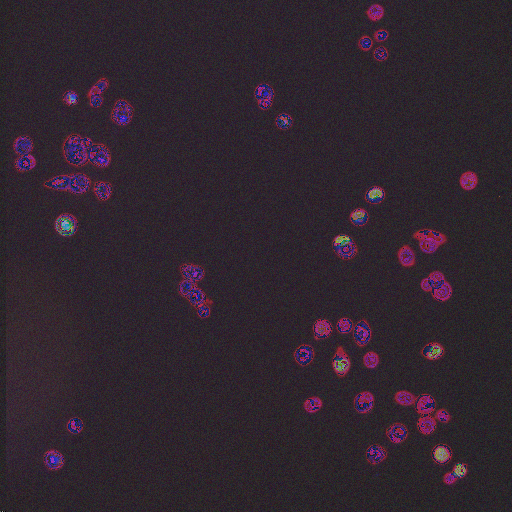
Large Spread |
Phospho-histone H3 dots |
Prometaphase |
Prophase |
Biological application
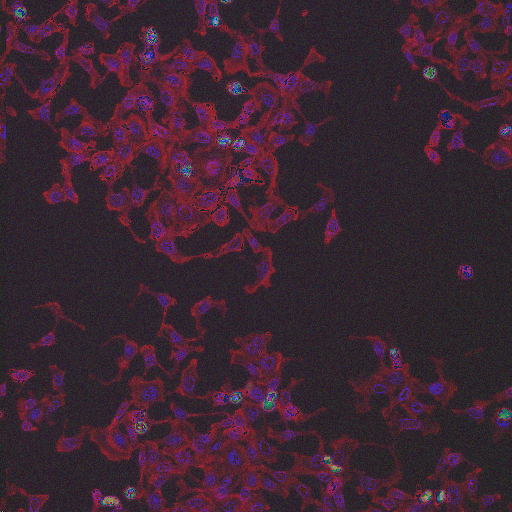
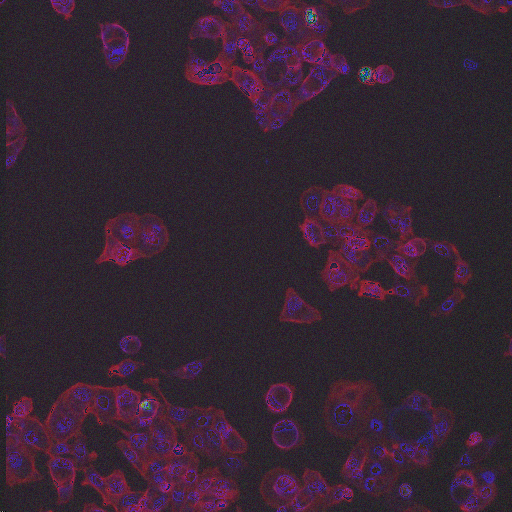
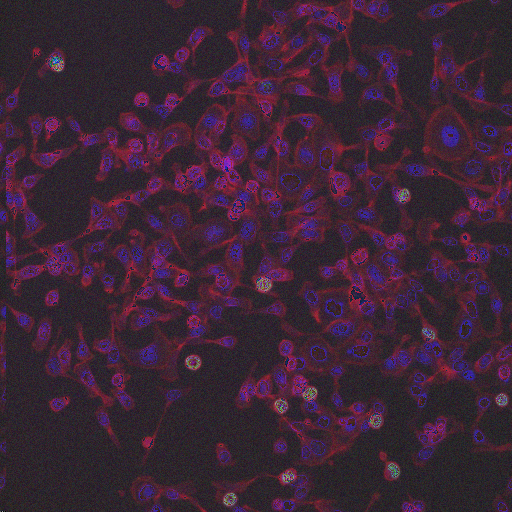
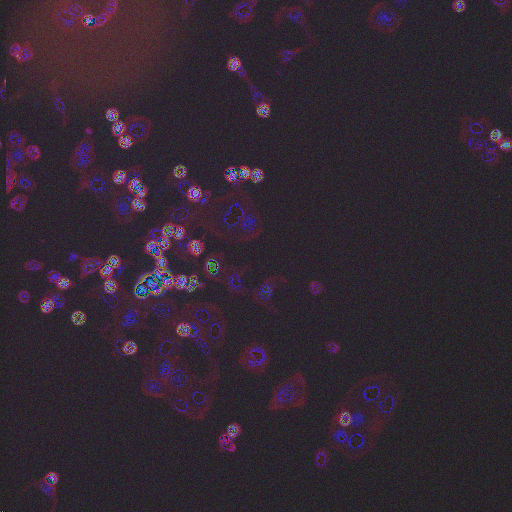
HT29 is a human colon-cancer cell line that has been widely used for the study of many normal and neoplastic processes. In order to obtain biologically relevant measurements of the cells, it is typically necessary to segment each nucleus and cell. Although image set BBBC008 can be used to quantify an algorithm's ability to segment nuclei and cells for this phenotype, there is always the concern with high-throughput experiments that some perturbants may lead to dramatically different morphologies, and that a segmentation algorithm may not be robust to such morphological variations.
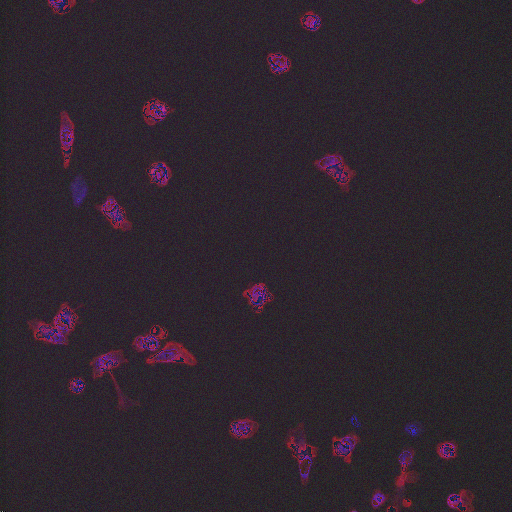
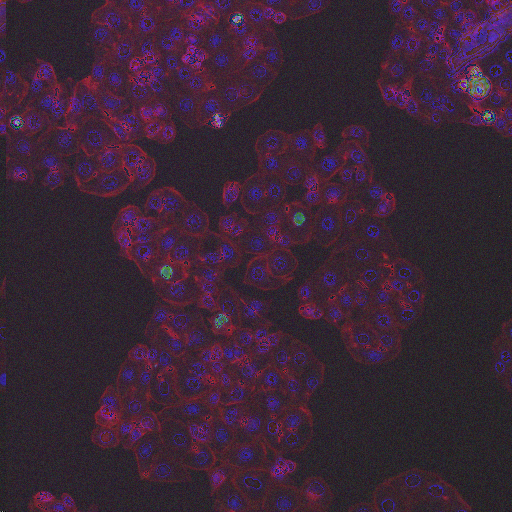
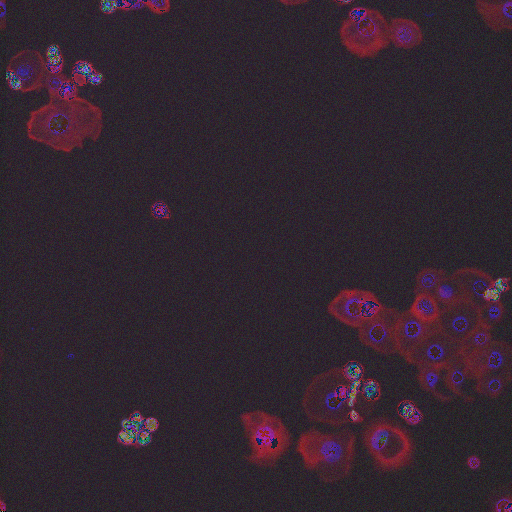
In order to form a basis for measuring a segmentation algorithm's robustness to variations in morphology, this image set contains four images enriched for each of 14 diverse morphologies of HT29 cells. The images are from Moffat et al.'s RNAi screen (BBBC017, and the enriched images were from the top-scoring sample for each of the 14 classifiers trained by Jones et al. (Proc Nat Acad Sci USA, 2009).
Images
The image set consists of 56 fields of view (4 from each of 14 samples). Because there are three channels, there are 168 image files. (The samples were stained with Hoechst 33342, pH3, and phalloidin. Hoechst 33342 is a DNA stain that labels the nucleus. Phospho-histone H3 indicates mitosis. Phalloidin labels actin, which is present in the cytoplasm.) The samples are the top-scoring sample from each of Jones et al.'s classifiers, as listed in the file SamplesScores.zip in their supplement. The files are in DIB format, as produced by the Cellomics ArrayScan instrument at the Whitehead–MIT Bioimaging Center. We recommend using Bio-Formats to read the DIB files. Each image is 512 x 512 pixels.
The filenames are of the form wellidx-channel.DIB, where wellidx is the five-digit well index (from Jones et al.'s supplement) and channel is either DNA, actin, or pH3, depending on the channel.
Ground truth

BBBC018_v1_outlines.zip (2.8 MB)
Wellidx 10779 has been excluded because the images were too blurred for the cells and nuclei to be segmented with confidence.
Recommended citation
"We used image set BBBC018v1 from the Broad Bioimage Benchmark Collection [Ljosa et al., Nature Methods, 2012]."
Copyright
 The BBBC018 images and ground truth are licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License by David Root and Anne Carpenter.
The BBBC018 images and ground truth are licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License by David Root and Anne Carpenter.