Accession number BBBC022 · Version 1
Example Images
 |
 |
 |
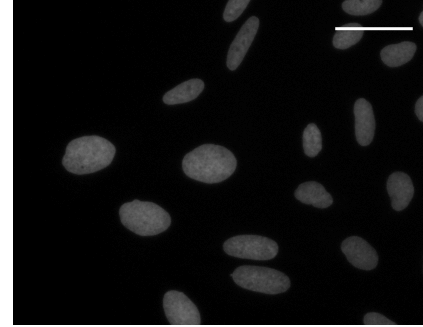
Hoechst 33342 staining |
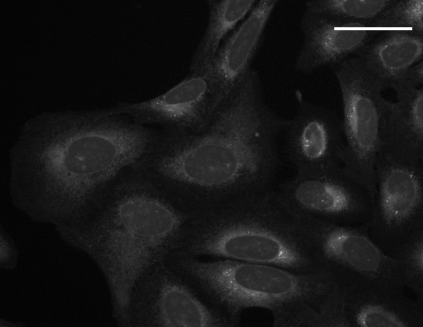
con A staining |
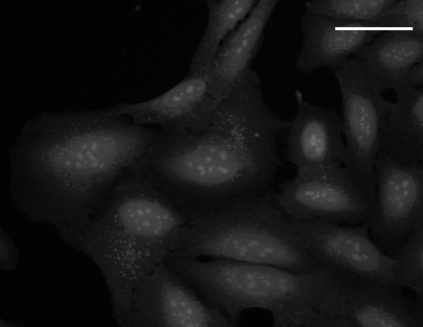
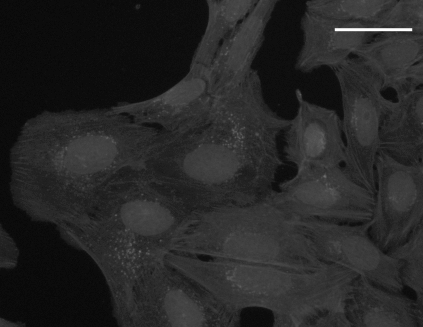
SYTO 14 staining |
 |
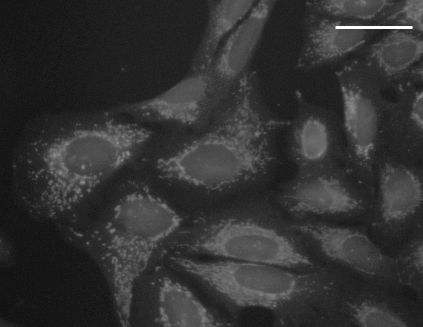 |
WGA + phalloidin staining |
MitoTracker Deep Red staining |
Description of the biological application
Phenotypic profiling attempts to summarize multiparametric, feature-based analysis of cellular phenotypes of each sample so that similarities between profiles reflect similarities between samples. This image set provides a basis for testing image-based profiling methods wrt. to their ability to distinguish the effects of small molecules.
Images
The images are of U2OS cells treated with each of 1600 known bioactive compounds and labeled with six labels that characterize seven organelles (the “Cell Painting” assay).
This pilot experiment consists of 20 plates. Each plate has 384 wells and each well has 9 fields of view for a total of 69,120 fields of view. Each field was imaged in five channels (detection wavelengths), and each channel is stored as a separate, grayscale image file, so there are 345,600 image files in 16-bit TIFF format. The images were taken at 20X magnification (0.656 μm/pixel).
The images are provided in 100 zip archive files, one for each combination of plate and channel. There is a list of the URLs of all 100 files to facilitate downloading the entire image set in batch.
Metadata
The file BBBC022_v1_image.csv (35 MB) contains the metadata in CSV format, with the following fields:
| Field name | Explanation |
|---|---|
| ImageNumber | a number between 1 and 69,120 identifying the field |
| Image_FileName_OrigER | name of image file for the con A channel |
| Image_FileName_OrigHoechst | name of image file for the Hoechst 33342 channel |
| Image_FileName_OrigMito | name of image file for the MitoTracker Deep Red channel |
| Image_FileName_OrigPh_golgi | name of image file for the WGA/phalloidin channel |
| Image_FileName_OrigSyto | name of image file for the SYTO 14 channel |
| Image_Metadata_ASSAY_WELL_ROLE | "compound" or "mock" |
| Image_Metadata_BROAD_ID | identifies the compound |
| Image_Metadata_CLSID | |
| Image_Metadata_CPD_MMOL_CONC | concentration of the compound |
| Image_Metadata_CPD_PLATE_MAP_NAME | name of plate map (stock plate) |
| Image_Metadata_CPD_SMILES | structure of the compound |
| Image_Metadata_CPD_WELL_POSITION | position of well on the plate |
| Image_Metadata_PlateID | name of plate |
| Image_Metadata_SOURCE_COMPOUND_NAME | compound name according to source |
| Image_Metadata_SOURCE_NAME | source of compound |
| Image_Metadata_Site | site number within well |
| Image_Metadata_Time | time the image was acquired (ms) |
| Image_PathName_OrigER | path to image file for the con A channel |
| Image_PathName_OrigHoechst | name of image file for the Hoechst 33342 channel |
| Image_PathName_OrigMito | path to image file for the MitoTracker Deep Red channel |
| Image_PathName_OrigPh_golgi | path to image file for the WGA/phalloidin channel |
| Image_PathName_OrigSyto | path to image file for the SYTO 14 channel |
Example rows from the file:

Nucleus segmentation annotations
Part of this dataset has been used for nucleus segmentation research, and a subset of 200 images have been selected for manual annotation of nucleus shapes. These annotations and more details about the segmentation problem can be found in the BBBC039 dataset.
Recommended citation
"We used image set BBBC022v1 [Gustafsdottir et al., PLOS ONE, 2013], available from the Broad Bioimage Benchmark Collection [Ljosa et al., Nature Methods, 2012]."
Copyright
 To the extent possible under law, Anne Carpenter has waived all copyright and related or neighboring rights to BBBC022v1. This work is published from: United States.
To the extent possible under law, Anne Carpenter has waived all copyright and related or neighboring rights to BBBC022v1. This work is published from: United States.