A dataset of images and morphological profiles of 30 000 small-molecule treatments using the Cell Painting assay
Accession number BBBC047 · Version 1
Example images
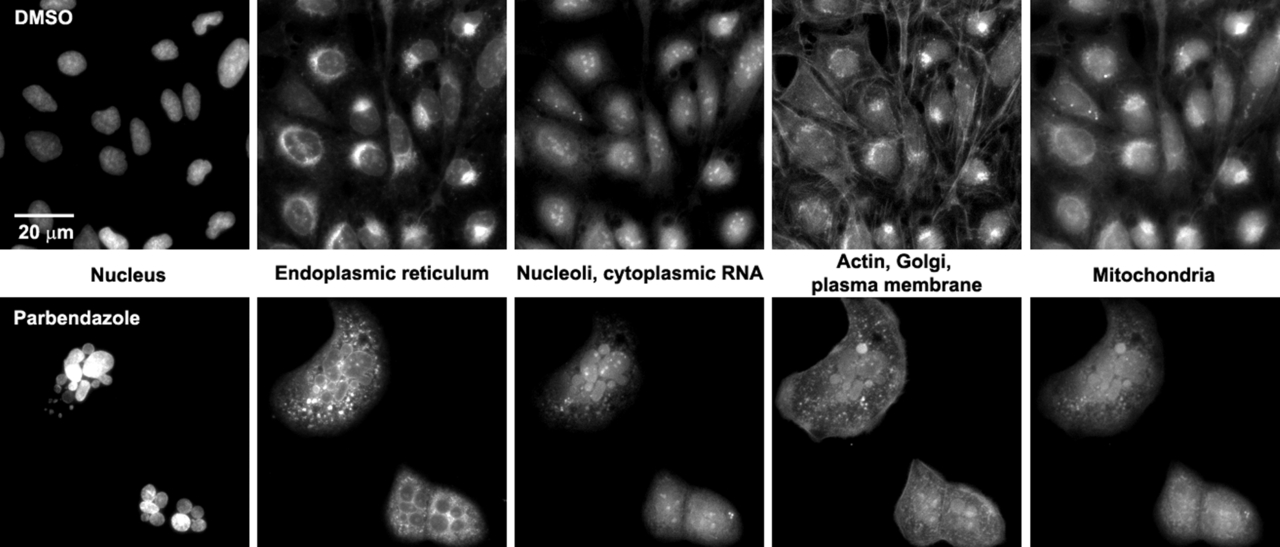
(Figure from the paper)
Biological application
This BBBC entry is a pointer to our previously published dataset. The abstract is copied below.
The images from the dataset are available on IDR.
The image-based profiles are available on the accompanying GigaDB web page.
The GitHub repo for the GigaDB entry is found here.
Background
Large-scale image sets acquired by automated microscopy of perturbed samples enable a detailed comparison of cell states induced by each perturbation, such as a small molecule from a diverse library. Highly multiplexed measurements of cellular morphology can be extracted from each image and subsequently mined for a number of applications.
Findings
This microscopy dataset includes 919 265 five-channel fields of view, representing 30 616 tested compounds, available at “The Cell Image Library” (CIL) repository. It also includes data files containing morphological features derived from each cell in each image, both at the single-cell level and population-averaged (i.e., per-well) level; the image analysis workflows that generated the morphological features are also provided. Quality-control metrics are provided as metadata, indicating fields of view that are out-of-focus or containing highly fluorescent material or debris. Lastly, chemical annotations are supplied for the compound treatments applied.
Conclusions
Because computational algorithms and methods for handling single-cell morphological measurements are not yet routine, the dataset serves as a useful resource for the wider scientific community applying morphological (image-based) profiling. The dataset can be mined for many purposes, including small-molecule library enrichment and chemical mechanism-of-action studies, such as target identification. Integration with genetically perturbed datasets could enable identification of small-molecule mimetics of particular disease- or gene-related phenotypes that could be useful as probes or potential starting points for development of future therapeutics.
Images
We used the Cell Painting assay. The images from the dataset are available on IDR
Ground truth 
Please see the paper for the details. The GitHub issue discussing details of this dataset is here https://github.com/broadinstitute/imaging-bbbc/issues/61 (Please post your questions in that issue thread).
For more information
Please contact Shantanu Singh of the Carpenter-Singh Lab at the Broad Institute of MIT and Harvard regarding this dataset.
Published results using this image set
See Google Scholar citations here.
Recommended citation
"We used image set BBBC047v1, available from the Broad Bioimage Benchmark Collection [Ljosa et al., Nature Methods, 2012]."
Copyright

The images and ground truth are licensed under a Creative Commons Attribution 3.0 Unported License (Commercial use allowed)